Full Service RNAseq
RNA Quality Control
Accurate evaluation of RNA quality and quantity is critical to produce high quality RNAseq data and is often the primary factor used to determine the optimal library prep methodology. RNA QC is included as part of all of library prep services and is also available as a stand alone service. RNA is assayed using the Agilent (AATI) Fragment Analyzer or Agilent Bioanalyzer to generate concentration measurement, RIN/RQN score, and/or DV200 metric.
RIN or RQN
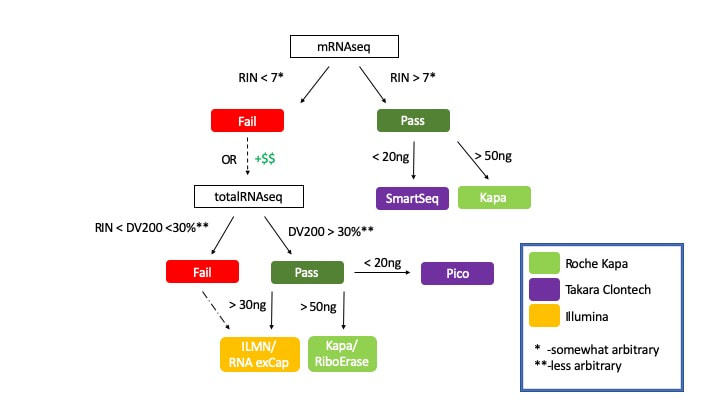
RNA Integrity Number (RIN) or RNA Quality Number (RQN) are similar metrics generated by the Bioanalyzer and Fragment Analyzer respectively. They are calculated based on the the relative intensity of the 18S and 28S ribosomal RNA electrophoretic peaks. Although somewhat arbitrary, we use a cutoff of 7 or more to define “high quality” RNA suitable for polyA enrichment library prep methods (mRNAseq). When dealing with degraded RNA, DV200 is a much more appropriate metric to make informed protocol decisions.
DV200: Fragment Distribution Value > 200nt
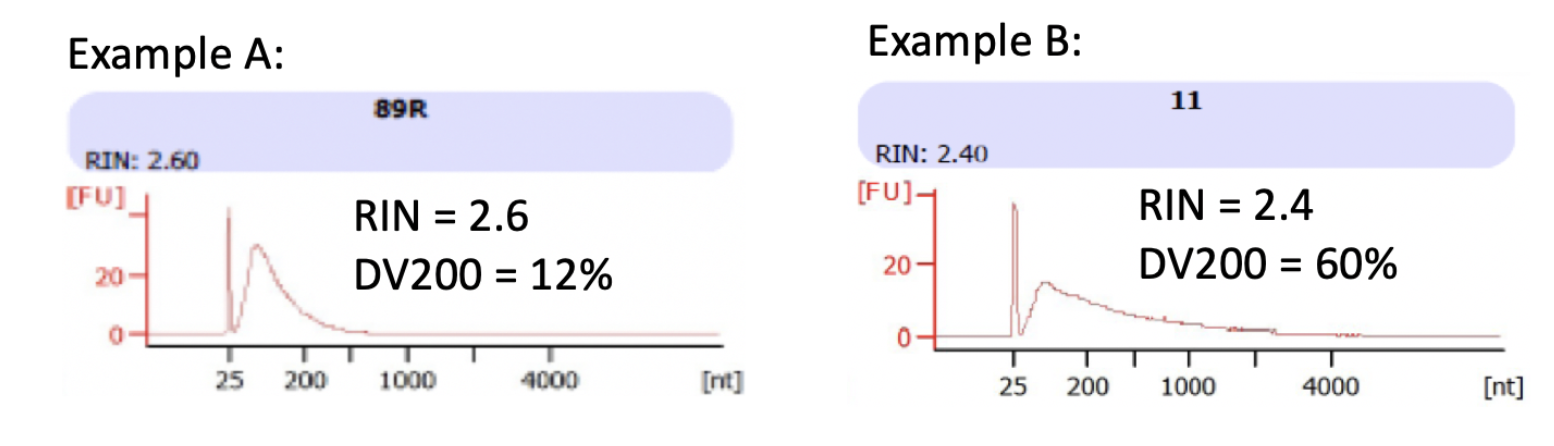
DV200 is the measure of the percentage of RNA that is greater than 200 nucleotides. This metric is more informative than RIN/RQN scores because it more accurately capture the extent of degradation. And most importantly, it informs the fraction of the RNA sample that can likely be converted into sequencing library. Samples may have similar RIN scores, but very different DV200s (see examples below).
RIN or RQN
RNA Integrity Number (RIN) or RNA Quality Number (RQN) are similar metrics generated by the Bioanalyzer and Fragment Analyzer respectively. They are calculated based on the the relative intensity of the 18S and 28S ribosomal RNA electrophoretic peaks. Although somewhat arbitrary, we use a cutoff of 7 or more to define “high quality” RNA suitable for polyA enrichment library prep methods (mRNAseq). When dealing with degraded RNA, DV200 is a much more appropriate metric to make informed protocol decisions.
DV200: Fragment Distribution Value > 200nt
DV200 is the measure of the percentage of RNA that is greater than 200 nucleotides. This metric is more informative than RIN/RQN scores because it more accurately capture the extent of degradation. And most importantly, it informs the fraction of the RNA sample that can likely be converted into sequencing library. Samples may have similar RIN scores, but very different DV200s (see examples below).
HOME | ABOUT US | SERVICES | SAMPLE SUBMISSION | DATA TRANSFER | PRICING | FAQ
Contact US: [email protected]
©2024 Molecular Biology Core Facilities at Dana-Farber Cancer Institute
Contact US: [email protected]
©2024 Molecular Biology Core Facilities at Dana-Farber Cancer Institute